| | Accueil | Annonces | Arts | Calendar | Cartes | Chat | Commentaires | Communautés | Coutumes | Culture | Délires | | |
| | Enregistrement | e-Souks | Forum | Fun | Galerie Photo | Genealogie | Harissatheque | Histoire | JRencontre | | |
| | Kandil | KesTuCherch | La bouffe | Links | Loterie | Medias | Musique | Objectif | Paracha | Plan du site | Portail | | |
| | Recherche | Religion | Sommaire | Sondages | Souvenirs | Telecom | Tunes Célčbres | Voyages | | |
|
|
A Y-CHROMOSOME PORTRAIT OF THE POPULATION OF JERBA (TUNISIA) |
A Y-CHROMOSOME PORTRAIT OF THE POPULATION OF JERBA (TUNISIA) TO ELUCIDATE ITS COMPLEX DEMOGRAPHIC HISTORY
ÉTUDE DE LA VARIABILITÉ DU CHROMOSOME Y DANS LA POPULATION DE JERBA (TUNISIE) AFIN D’ÉLUCIDER SON HISTOIRE DÉMOGRAPHIQUE
Franz MANNI 1, Pascal LEONARDI 1, Étienne PATIN 1, Alain BERREBi 2, Houssein KHODJET EL KHIL 3, Karl SKORECKI 4, Dror ROSENGARTEN 4, Hassan ROUBA 5, Évelyne HEYER 1, Marc FELLOUS 6
ABSTRACT
The Island of Jerba has a long demographic history that is related to its central position in the Mediterranean Sea. The island was in contact with different civilizations and its population is heterogeneous (Arabs, Berbers, black Africans and Jews).
We sampled 127 males (Jewish sample N = 32; Berber sample N = 46; Arab sample N = 47) in order to identify the Y-chromosome variability of the population according to 10 Unique Event Polymorphisms (UEPs): SRY-2627;
SRY-10831a; SRY-4064; 92R7; Tat; YAP; M2; LLY22g; M9; 12f2q. We also sampled the Black group living in Jerba but typing ambiguities forced us to exclude it from further analyses.
The results suggest a very low degree of genetic differentiation between Jerban Arabs and Berbers, who can be considered genetically, as belonging to the same group. In contrast, pairwise FST measures show that the genetic distances between the Jewish sample and the other ethnic groups are the highest in the island. This datum implies a low level of gene flow between the different communities that reside in Jerba, with the exception of Arabs and Berbers. Since geographic isolation plays no role, the different allelic profiles of the three ethnic groups are related to their different geographic origins, which are likely to have been maintained by the cultural differences existing between Arabs, Berbers and Jews.
By comparing the observed haplogroup profiles with 19 reference populations located around the Mediterranean basin, we confirm a North African origin for the Berber and Arab sample and a Middle Eastern ancestral population for Jerban Jews.
Keywords: Jerba, Y-chromosome, Unique Event Polymorphisms (UEPs).
Bulletins et Mémoires de la Société d’Anthropologie de Paris, n.s., t. 17, 2005, 1-2, p.
1. Équipe de Génétique des Populations, Département Hommes, Natures, Sociétes, Musée de l’Homme, MNHN, 17 place du Trocadéro et du 11 novembre, 75016 Paris, France, e-mail: manni@mnhn.fr
2. Hematology Institute, Kaplan Medical Center, Rehovot, Israel.
3. Laboratoire de Génétique Moléculaire, Immunologie et Biotechnologie, Faculté des Sciences, Tunis, Tunisie.
4. Technion, Israel Institute of Technology, Haifa, Israel.
5. Institut Pasteur, Casablanca, Morocco.
6. Inserm E0021, Hôpital Cochin, 75014 Paris, France.
2 F. MANNI, P. LEONARDI, É. PATIN, A. BERREBI, H. KHODJET EL KHIL,
K. SKORECKI, D. ROSENGARTEN, H. ROUBA, É. HEYER, M. FELLOUS
RÉSUMÉ
L’histoire démographique de l’île de Jerba est trčs ancienne en raison de sa position centrale dans le bassin de la mer Méditerranée, c’est la raison pour laquelle plusieurs populations se sont succédées sur l’île dont la population est, de nos jours, hétérogčne (Arabes, Berbčres, Noirs, Juifs).
Afin d’étudier la variabilité du chromosome Y de la population de Jerba nous avons analysé 10 polymorphismes, SRY-2627 ; SRY-10831a ; SRY-4064 ; 92R7 ; Tat ; YAP ; M2 ; LLY22g ; M9 ; 12f2q sur un échantillon de 127 individus (Juifs N = 32 ; Berbčres N = 46 ; Arabes N = 47). Nous avions aussi échantillonné la population noire de Jerba mais des ambiguďtés, lors du génotypage, nous ont forcé ŕ exclure l’échantillon de l’article.
Les résultats indiquent une trčs faible différenciation génétique entre les échantillons arabe et berbčre qui peuvent ętre considérés, d’un point de vue génétique, comme une seule population. Inversement, les mesures de FST montrent que les distances entre l’échantillon juif et les autres groupes ethniques de l’île sont considérables. Ce résultat implique un faible niveau d’échange génétique entre les communautés qui habitent l’île, ŕ la seule exception des Arabes et des Berbčres.
Comme l’isolement géographique est inexistant ŕ Jerba, les différences des profils alléliques entre les trois groupes s’expliquent par une provenance géographique différente et une barričre culturelle relative concernant Arabes, Berbčres et Juifs.
La comparaison entre la composition haplotypique de Jerba et celle de 19 populations de référence, localisées autour du bassin de la mer Méditerranée, montre la provenance nord-africaine des Berbčres et des Arabes de Jerba. De la męme façon, l’échantillon juif vient d’une population mčre localisée au Moyen-Orient.
Mots-clés : Jerba, chromosome Y, Unique Event Polymorphisms (UEPs)
.I was driven thence by foul winds for a space of nine days upon the sea, but on the tenth day we reached the land of the Lotus-eater, who live on a food that comes from a kind of flower. Here we landed to take in fresh water, and our crews got their mid-day meal on the shore near the ships. When they had eaten and drunk I sent two of my company to see what manner of men the people of the place might be…
Homer, The Odyssey, Book IX (Translation by S. Butler)
INTRODUCTION
The island of Jerba, located in south-eastern Tunisia, represents an interesting case-study for human population genetics studies because a large amount of genetic data concerning the island is becoming available. Published studies portray its variability in details ranging from autosomal markers, classic blood markers (Gherib 1962; Ranque et al. 1964; Mourali 2000; Yacoubi Loveslati
et al. 2001) and uniparental markers such as mitochondria (Loveslati 1999) and Y-chromosome (Khodjet El Khil et al. 2001, in press). It is known that different markers can mirror different demographic histories and that possible discrepancies can relate to the different patterns of genetic transmission existing between recombining and no-recombining markers. Interestingly, the analysis of non-recombining markers can reveal differences between the demographic history of females and males, which can indicate a different history of migrations as in patrilocality or matrilocality (Seielstad et al. 1998; Hammer et al.2001; Oota et al. 2002; Kayser et al. 2003).
A Y-CHROMOSOME PORTRAIT OF THE POPULATION OF JERBA (TUNISIA) 3
Besides the value of a comparative approach between different markers, an additional importance of the island of Jerba is related to its location in North Africa.
This article follows a previous study concerning Egypt (Manni et al. 2002) within the wider framework of a project on the genetic patterns of variation in North Africa. The study of the north-eastern part of Africa is likely to provide a deeper understanding of i) remote human history as in the farming/language dispersals hypothesis (Krings et al. 1999; Manni et al. 2002; Lucotte, Mercier 2003) as well as ii) more recent migrations around the Mediterranean basin. We have shown that a) the genetic history of Egypt is very complex with sub-Saharan, Middle Eastern and European contributions to its genetic pool and confirmed that b) there are no genetic signatures of admixtures between NW Africa and southern Europe (Manni et al. 2002).
Despite the particular value of North Africa, only a few studies have defined the genetic background prevailing in modern populations from this region. We focus here on Jerba, which on a smaller scale appears to reflect this complexity. There are four distinct human groups which have cohabited on the island for centuries, and who identify themselves as having a different ethnic origin: Arabs, Berbers, Blacks and Jews. Moreover, since Jerba is an island, possible local admixtures are likely to be traced back more successfully because of the absence of geographic continuity with surrounding populations.
According to North African history, Berbers are believed to have their ancestors among Capsians (7000- 5000 B.C.) (Cavalli-Sforza et al. 1994), and several other historical records can be useful in tracing back the demographic history of this island, thus providing a strong historical background for interpretation of the genetic variation of the samples (Mourali 2000). The first Arab settlement on the island occurred in the 7th century A.D. when Islam and the Arabic language were imposed upon the Berbers. It has been proposed that the black Africans were descendants of Sudanese or Nigerians slaves who came in the 18th century A.D. (Tlatli 1967). The origin of the Jews appears to be more complex since differentmigrations have been proposed, one represented by Jews that migrated to the island after the destruction of the Temple in Jerusalem in the 6th century B.C., a second one during the 15th century A.D. Furthermore, additional migrations were suggested (Auctores). This picture is further complicated by the fact that between the 9th century B.C. and the 19th century A.D., Jerba was a pivotal area in the migration routes of several populations:
Phoenicians, Romans, Byzantines, Bedouins, Germans, Sicilians, Turkish and French (Tlatli 1967).
Currently, Berbers, Blacks and Jews live in different areas of Jerba, whereas only Arabs are distributed over the whole island. The main language spoken by all islander groups is Arabic, even though Berbers also speak a dialect called “Chelha” that belongs to Shilha (a cover term for Berber languages in Marocco and Tunisia). Religious differences may have played a role in subdividing populations; Arabs and Blacks are orthodox Muslims, Berbers are heterodox Muslims while the Jewish group follows Judaism.
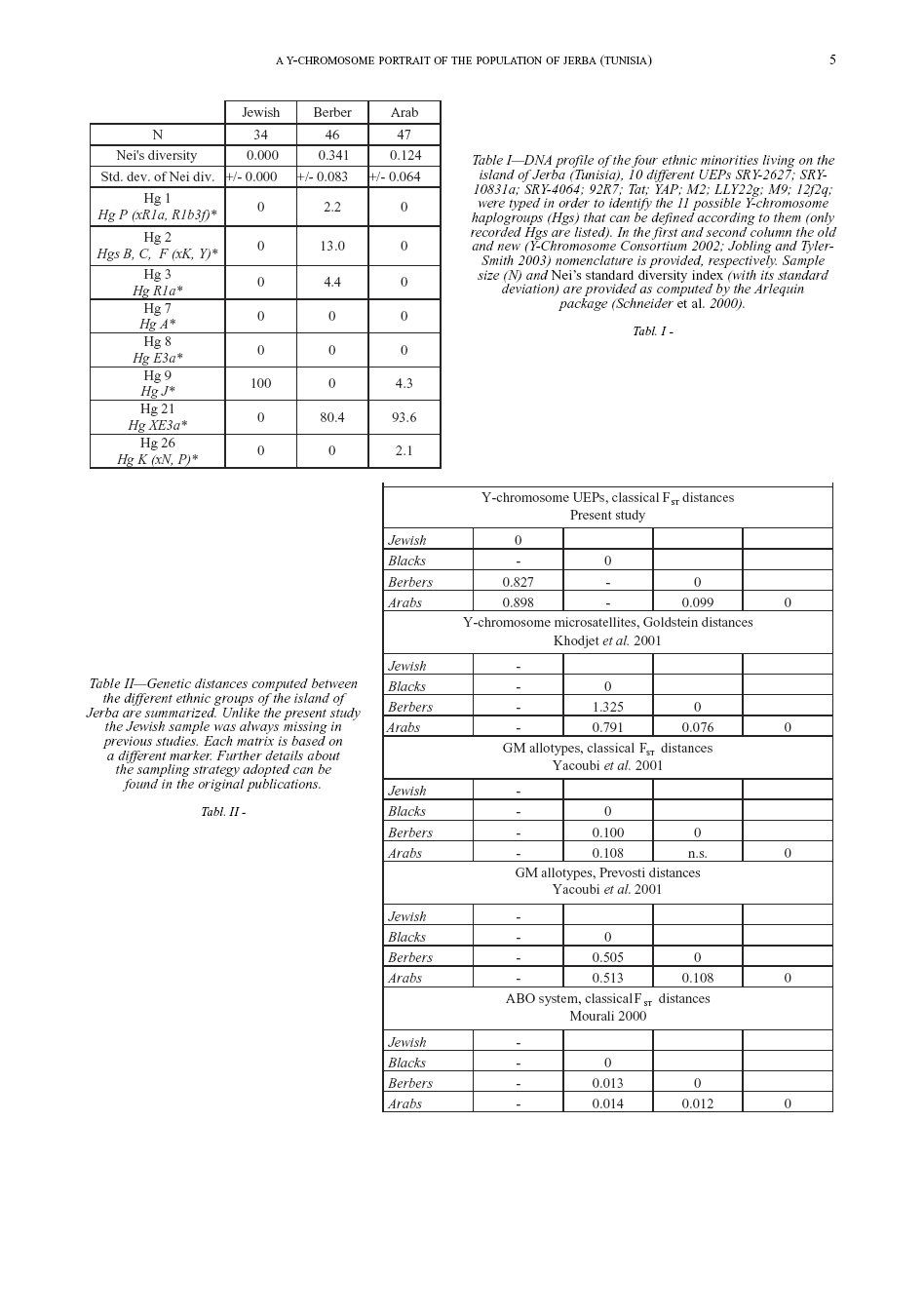
To portray the genetic differences and variability of the present Jerban population, we present the allelic profile of eleven specific Unique Event Polymorphisms (UEPs) meaning that each mutation defines a group of evolutionary related Y-chromosomes (haplogroups). This study is intended to provide a better understanding of the origin of the four ethnic groups living in Jerba.
MATERIALS AND METHODS
Sample
Genetic analyses were performed on a sample of 127 Y-chromosomes representing 47 Jerban Arabs, 46 Jerban Berbers and 34 Jerban Jews. Since a large majority of Jerban Jews migrated to Israel in 1948 A.D., we sampled such recent immigrants from Jerba to the Israeli towns of Safed, Haifa, Ashdod, Gedera, Ashkelon, Brekhia, Eitan and Tlamim. Only Jews with a Jerban ancestry of at least four generations were included in the study. This criterium also applies the DNA donors belonging to the other ethnic groups. Overall, our sample represents almost 1‰ of the total population of 114,170 individuals. Each Y-chromosome comes from different unrelated families. Fully informed consent was obtained from all participants in this study.
Y-chromosome biallelic polymorphisms typing DNA was extracted from whole blood by means of a standard phenol/chlorophorm protocol. Subsequent purified DNA was analyzed in order to identify the 11 possible Y-chromosome haplogroups (Hgs) defined by 10 UEPs (Unique Event Polymorphisms) (Y-Chromosome Consortium 2002; Jobling, Tyler-Smith 2003). The markers used are the as follows: SRY-2627
4 F. MANNI, P. LEONARDI, É. PATIN, A. BERREBI, H. KHODJET EL KHIL, K. SKORECKI, D. ROSENGARTEN, H. ROUBA, É. HEYER, M. FELLOUS
(Veitia et al. 1997), SRY-10831a (Kwok et al. 1996), SRY-4064 (Whitfield et al. 1995), 92R7 (Mathias et al. 1994), Tat (Zerjal et al. 1997), YAP (Hammer, Horai 1995), M2 (Seielstad et al. 1994), LLY22g (E. Righetti, C. Tyler-Smith—unpublished), M9 (Underhill et al. 1997) and 12f2q (Casanova et al. 1985). The typing was carried out as described in Rosser et al. (2000). The deep-rooting markers SRY-10831a, M9, 92R7, YAP and 12f2q were typed in all samples and, in some cases, the remaining markers were typed hierarchically; e.g. SRY-4064 and M2 only on YAP+ chromosomes, LLY22g and Tat only on those chromosomes which are M9 derived and 92R7 ancestral, and SRY-2627 only on those chromosomes which are 92R7 derived.
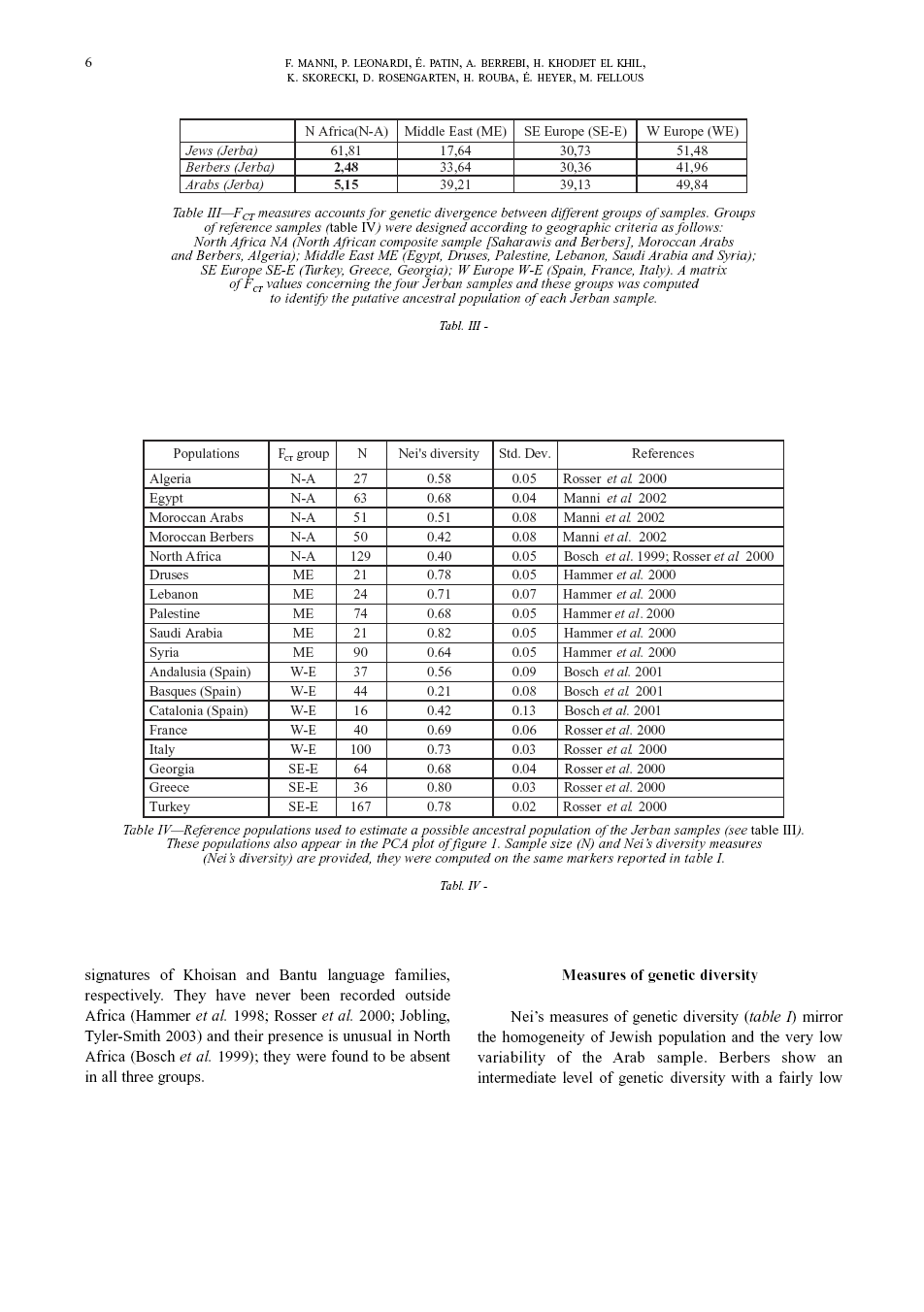
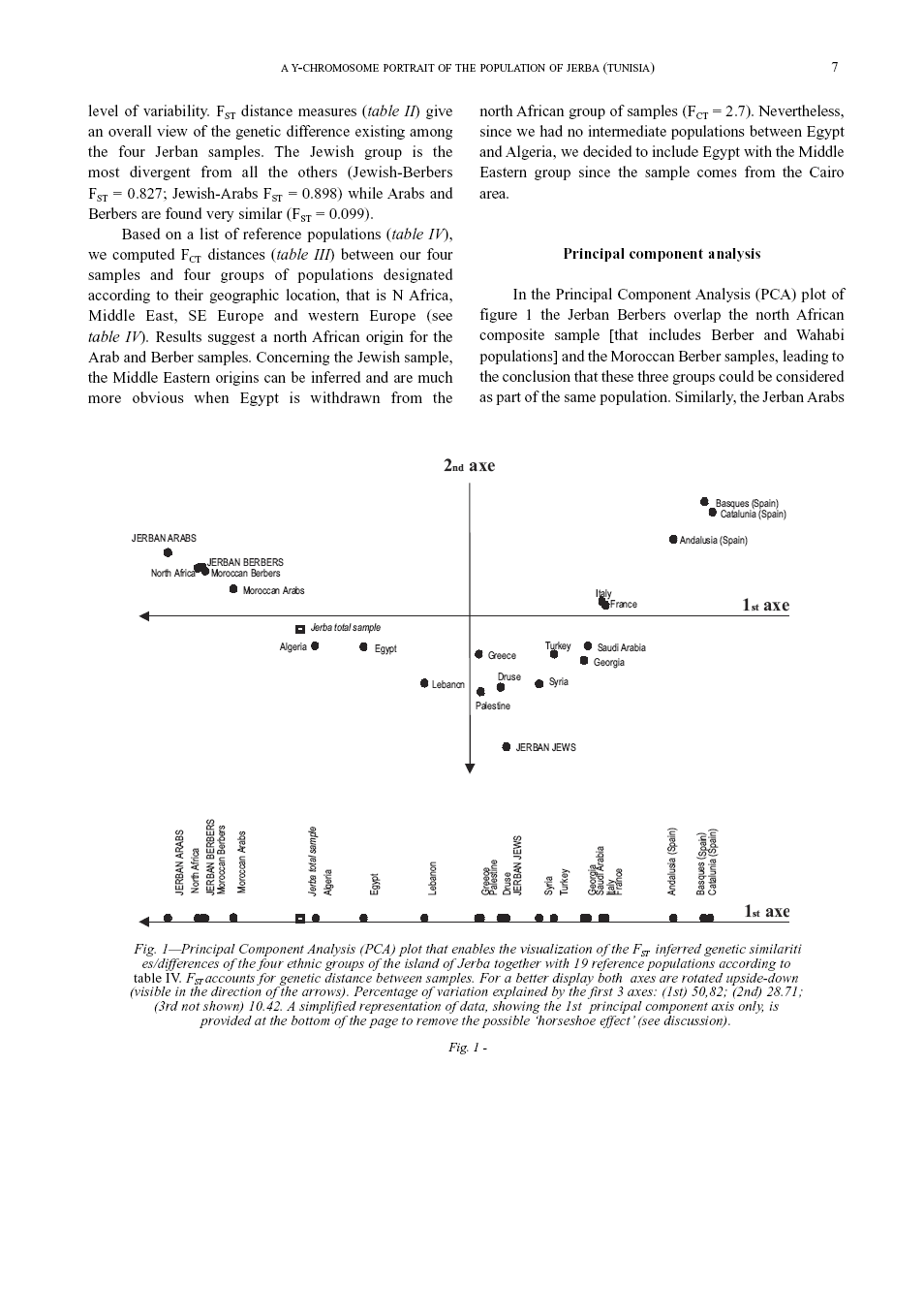
Measures of genetic diversity and differentiation Two genetic diversity estimators were computed by means of the Arlequin version vs. 2.1 software (Schneider et al. 2000). Intra-population diversity was measured through the gene diversity index (Nei 1987) as shown in table I. Inter-population diversity was calculated through conventional F-statistics (Wright 1965). Two measures of differentiation are provided: FST accounts for genetic divergence between samples (table II); FCT accounts for genetic divergence between different groups of samples (table III). A group of reference samples (table III, IV) was designated according to geographic criteria as follows: NW-Africa (North African composite sample [includes different Berber and Wahabi populations], Morrocan Arabs and Berbers, Algerians); Middle East (Egyptians, Druses, Palestinians, Lebanese, Saudi Arabians and Syrians); SE-Europe (Turks, Greeks, Georgians); western Europe (Spanish, French, Italian). A matrix of FCT values concerning the four Jerban samples and these groups was computed to identify the putative mother population of each Jerban sample (table III).
Multivariate analysis
Principal component analysis (PCA) was applied to the data to graphically identify possible patterns of genetic similarity between the four Jerban ethnic groups and a reference sample of 19 populations around the Mediterranean basin (table IV). The PCA method involves a mathematical procedure that transforms a number of (possibly) correlated variables into a (smaller) number of uncorrelated variables called principal components. The first principal component accounts for as much of the variability in the data as possible, and each succeeding component accounts for as much of the remaining variability as possible. Multidimensional relationships between items can be seen in a bivariate plot (if two or three axes are plotted one against each other). The analysis was performed with the Excel applet GenAlEx by Peakall and Smouse (2001) freely available at the following Internet address: http://www.anu.edu.au/BoZo/genAlEx
RESULTS
Y-chromosome biallelic profiles
Allelic profiles of the 11 Y-chromosome lineages in the total sample of 165 Jerban Y-chromosomes are shown in table I. The first result concerns the total homogeneity of the Jewish group where all the 34 typed individuals share the same haplogroup Hg J that is also present, at a very low frequency (4.3%), in the Arab group. This haplogroup presents its highest frequencies in the Fertile Crescent and has been suggested to be a genetic signature of migrations from the Middle East in Neolithic and post-Neolithic times (Semino et al. 1996, 2004; Rosser et al.
2000; Quintana-Murci et al. 2001; Arredi et al. 2004;
Di Giacomo et al. 2004).
The only haplogroup spread between the remaining two ethnic groups is the 21. High frequencies of this lineage are recorded in the Jerban Berbers (80.4%) and Arabs (93.6%). This haplogroup shows its highest frequencies in North Africa. Hg P(xR1a, R1b3f) lineage has been found predominantly in Europe, showing increasing frequencies from the Middle East to northwestern Europe (Semino et al. 1996; Rosser et al. 2000) and was found at a very low frequency in Berbers (2.2%).
Hgs B, C and F (xK, J) is found at significant frequency in Berbers (13.2%) but are absent in Arabs. However, little information can be deduced from this latter lineage since it is present in both Europe and Asia and does not show any informative geographic variation in the Eurasian landscape. We recorded the presence of Hg K (xN, P) in a single individual in the Arab sample. Given the little information conveyed by a single Y-chromosome we will not discuss it further in this context. Hg A and Hg E3a are found at high frequency in sub-Saharan African lineages and they have been suggested to be genetic
A Y-CHROMOSOME PORTRAIT OF THE POPULATION OF JERBA (TUNISIA) 5





| | Accueil | Annonces | Arts | Calendar | Cartes | Chat | Commentaires | Communautés | Coutumes | Culture | Délires | |
| | Enregistrement | e-Souks | Forum | Fun | Galerie Photo | Genealogie | Harissatheque | Histoire | JRencontre | |
| | Kandil | KesTuCherch | La bouffe | Links | Loterie | Medias | Musique | Objectif | Paracha | Plan du site | Portail | |
| | Recherche | Religion | Sommaire | Sondages | Souvenirs | Telecom | Tunes Célčbres | Voyages | |
Pour toutes informations, critiques et commentaires, envoyez un émail a :